| Discrete Dynamics Lab |
Update May 2016ddlabz04 xx This version corresponds exactly to the2nd Edition of Exploring Discrete Dynamics (EDD2)) (2016) -- the DDlab Manual -- available in preview.
May 2016 updates (on the
Sept 2015 release)
include the following new features, as well as improvements, bug fixes, and coordination between
the software and EDD2...
|

Resetting the transient scale for a single basin of attraction, v2k3 rcode (dec)30, $n$=12, seed=(hex)0baa --- a chain rule with long transients showing just a segment of the attractor (period 102) with a state highlighted. |
xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxx DDLab has been updated at regular intervals since its release in 1995. Its precursor was the Atlas software included on diskette inside the back cover of "The Global Dynamics of Cellular Automata" 1992. For a list and download of this and older versions click here.
Below are links to previous updates, |
|
EDD2#21.10, #21.11: Saving/loading seeds in ASCII and
Golly format. DDLab has its own efficient binary seed encoding (.eed files EDD2#21.9) but for interchanging a seed between DDLab and alternative software such as MatLab, the seed can also be saved/loaded in a simple ASCII encoding (EDD2#21.10) --- the first byte is the value-range v, then a byte for each value at index 0 to $n$-1.
| 
Golly is software in the game-of-Life community.
Golly's pattern (.rle} file format is an ASCII script following
``run-length'' encoding, For 2d binary patterns only, DDlab can
duplicate this encoding in its own (.gly) file, but omits Golly's
preamble in the top line. DDLab does not use or need the preamble
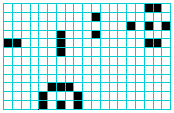
because the $x,y$ size is deduced from the script itself -- for
example, | x = 19, y = 12, rule = x-rule-pre 16b2o$10bo$14bobobo$6bo3bo$2o4bo9b2o$6bo4$5b3o$4bo3bo$4bobobo! encodes this 19x12 seed pattern on the left. To convert Golly's .gly to DDLab's .rle, a Golly preamble may be added and the file renamed. To convert .rle to .gly the Golly preamble is removed and the file renamed. | |
Return to the
Discrete Dynamics Lab home page.
Last modified: May 2016